SCIENCE
Engineered Living Materials (ELMs)
BIOFABRICATION
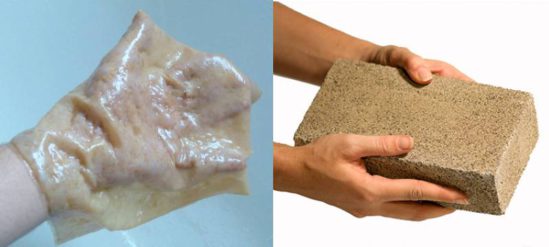
Microbially-synthesized nanocellulose (left) and biocement (right) show the promising future of living materials
(left image: Ahmed Zoheir, right: BioMasson)
Engineered living materials (ELMs) is an emerging field, in which living cells either create the material or regulate
its functionality. This unique approach utilizes microbial systems not just to produce useful raw materials as we used to, but interestingly to drive and control their assemblies into programmable patterns, and to architect novel materials. Such materials bear unique "living" features such as self-assembly, -regeneration and -repair. I am generally focused on developing novel Biofilm-Fabricated Engineered Living Materials (BioFELM) with the following interfaces:
- BioFELM»Catalysis -interface with biocatalysis
- BioFELM»Material -interface with materials science
- BioFELM»BuiltEn -interface with the built environment and architecture
- BioFELM»Design -interface with biodesign and art
As an example, bacterial cellulose (BC) is a rising star among ELMs. Due to its unique nanostructure, crystallinity, high water retention capacity as well as other mechanical properties, it has versatile potential applications such as wound dressing, tissue regeneration, drug delivery, filtration, actuation, and as leather substitute. I am interested in exploring novel BC applications, as well as engineering enhanced production strains and co-cultures with special focus on biofilms.
Microbially induced calcite precipitation (MICP) is another example for the use of ELMs in the built environment. MICP utilizes microorganisms' metabolic machinery and cell surface features to mineralize and nucleate inorganic calcite crystals. This technology can be used to produce living building materials with neutral or negative carbon fingerprint. Examples are the production of biocement, self-healing concrete as well as soil stabilization, dust control and surface protection. I am adopting the calcifying strains for new MICP applications, and exploring alternative MICP metabolic pathways.
Adaptive Laboratory Evolution (ALE)
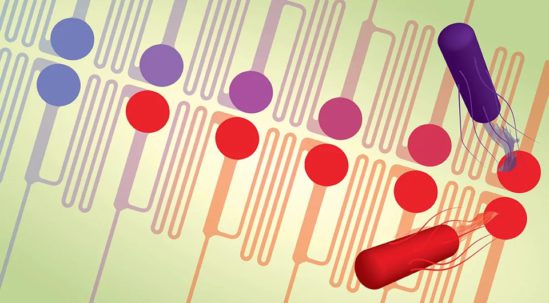
Evolution-on-a-chip
evo.S chip drives fast adaptation of microbial systems under continuous cultivation.
It differentiates between complex "persistent" and "resistant" phenotypes.
(Graphic: Ahmed Zoheir)

Microbes can be triggered to gain novel traits or to enhance their inherited characteristics, in an artificial process termed Adaptive Laboratory Evolution (ALE). ALE is an essential methodology answering fundamental questions ranging from the evolution of life and its underlying mechanisms to the adaptation of microbial populations to their environments. Especially in biotechnological production, ALE has been employed to improve yields or to enhance the organisms' tolerance to toxic conditions.
However, the common ALE techniques are either laborious or involve expensive equipment. Furthermore, they mainly employ well-mixed planktonic populations in a defined chemical environment and are thus paying less attention to the role of biofilm communities and spatial heterogeneity.
To address these limitations, a novel evolution-on-a chip system, dubbed "evo.S", has been developed that applies precise chemical gradients in consecutive microenvironments to enable miniaturization of microbial ALE. The evo.S chip has been employed to rapidly evolve E. coil resistance to the antibiotics rifampicin and nalidixic acid, encoded by newly discovered mutations. Adaptation on evo.S revealed the critical influence of the gradient profile on the adaptability of E. coli based on either phenotypic persistence or genetic resistance. Furthermore, the evo.S chip could be exploited for the development of enhanced industrial producer strains in miniaturized, highly parallel and cost-effective procedures.
Microbial Stress Response
STRUCTURES OF STRESS
A micrograph demonstrating bacteria under stress, reported by the RGB-S Reporter.
(Image: Ahmed Zoheir, IBG-1)
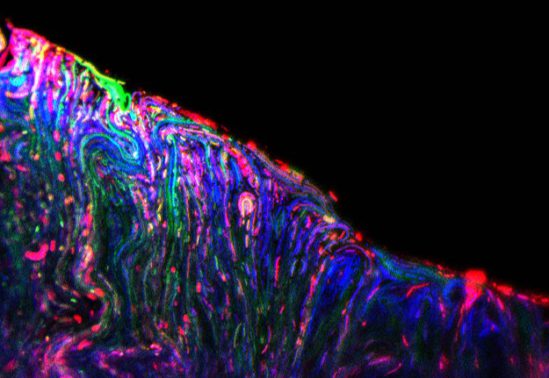
In their natural habitats as well as in biotechnological settings, bacteria are vulnerable to environmental fluctuations that often involve conditions causing cellular stress. In order to survive stress, specific response mechanisms have been evolved, which help bacteria to withstand such stressing threats.
The currently available stress-reporting systems lack the ability to report multimodal stress response in living cells with high spatiotemporal resolution. In one project, I have been able to develop the "RGB-S Reporter" - a novel multi-stress reporter system that enables simultaneous and independent sensing of bacterial response to genotoxicity, cytotoxicity and physiological stress. Using the state-of-the-art synthetic biology principles, this novel tool has been designed for precise reporting of cellular status in planktonic as well as biofilm populations, in normal cultivation devices as well as in modern microfluidic and micro-droplet systems.
Such kind of multi-stress biosensors enable the acquisition of high-content data to facilitate the fast, robust and comprehensive analysis of bacterial response to transient and permanent environmental changes. Such bio-based reporting systems are of great value not only for fundamental research in the context of stress heterogeneity within free-cells and biofilms, but also for practical applications such as stress-emergence of antibiotic resistance, assessment of environmental contamination and optimization of stress in biotechnological production processes.
Microfluidics & Productive Biofilms: UNDER CONSTRUCTION
Publications
JOURNAL ARTICLES
- Abdel-Gawwad, H.A., Saleh, A.A., Shoukry, H., Radwan, A.A., Abd Elrahman, M., & Zoheir, A.E. (2025). Biomediated Recycling of Aerated Concrete Waste into Thermally-insulating Living Building Materials. Chemical Engineering Journal, 166254. doi: 10.1016/j.cej.2025.166254.
- Zoheir A.E., Stolle C., Rabe K.S. Microfluidics for Adaptation of Microorganisms to Stress - Design and Application. Applied Microbiology and Biotechnology, 2024, 108(1), 162. doi: 10.1007/s00253-024-13011-x.
- Zoheir A.E., Meisch L., Sobol, M.S., Ordoñez-Rueda, D., Kaster, A.-K., Niemeyer C.M., Rabe K.S. A Three-colour Stress Biosensor Reveals Multimodal Response in Single Cells and Spatiotemporal Dynamics of Biofilms. npj Biofilms Microbiomes, 2023, 9, 57. doi: 10.1038/s41522-023-00424-1.
- Zoheir A.E., Meisch L., Martín M.V., Bickmann C., Kiselev A., Lenk F., Kaster A.-K., Rabe K.S., Niemeyer C.M. Macroporous Silicone Chips for Decoding Microbial Dark Matter in Environmental Microbiomes. ACS Applied Materials & Interfaces, 2022, 14, 44, 49592–49603. dio: 10.1021/acsami.2c15470.
- Zoheir A.E., Sobol, M.S., Ordoñez-Rueda, D., Kaster, A.-K., Niemeyer C.M., Rabe K.S. A Genetically-encoded Three-Colour Stress Biosensor Reveals Multimodal Response at Single Cell Level and Spatiotemporal Dynamics of Biofilms. bioRxiv, 2022, doi: 10.1101/2022.09.23.509207.
- Delavault A., Zoheir A.E., Muller D., Hollenbach R., Rabe K.S., Ochsenreither K., Rudat J., Syldatk C. Enhanced Bioactivity of Tailor-made Glycolipid Enriched Manuka Honey. International Journal of Molecular Sciences, 2022, 23, 12031.
- Bitterwolf P., Zoheir A.E., Hertel J., Kröll S., Rabe K.S., Niemeyer C.M. Intracellular Assembly of Interacting Enzymes Yields Highly-Active Nanoparticles for Flow Biocatalysis. Chemistry - A European Journal, 2022, 28.66. doi: 10.1002/chem.202202157.
- Lemke P, Zoheir A.E., Rabe K.S., Niemeyer C.M. Microfluidic Cultivation and Analysis of Productive Biofilms. Biotechnology and Bioengineering, 2021, 118(10), 3860-3870.
- Zoheir A.E., Späth G.P., Niemeyer C.M., Rabe K.S. A Microfluidic Evolution-on-a-Chip Reveals New Mutations that Cause Antibiotic Resistance. Small, 2021, 17(10), 2007166.
- Grösche M., Zoheir A.E., Stegmaier J., Mikut R., Mager D., Korvink J.G., Rabe K.S., Niemeyer C.M. Microfluidic Chips for Life Sciences—A Comparison of Low Entry Manufacturing Technologies. Small, 2019, 15, 1901956.
- Hammad I.A., Talkhan F.N., Zoheir A.E. Urease Activity and Induction of Calcium Carbonate Precipitation by Sporosarcina pasteurii NCIMB 8841. Journal of Applied Sciences Research, 2013, 9(3), 1525-1533.
ORAL PRESENTATIONS
- Zoheir A.E. Harnessing Microbial Biodiversity in Biotechnology: Microfluidics as Next-Generation Tools. Cairo Science Forum 23 / Humboldt Kolleg. 07-08.02.2023, Cairo, Egypt.
- Zoheir A.E. and Rabe K.S. Combination of Stress Sensors and Microfluidic Technologies for Accelerated Bacterial Adaptive Evolution. Retreat 2019 of the Biointerfaces International Graduate School (BIF-IGS). 11-13.05.2019, Trier, Germany.
- Zoheir A.E. and Rabe K.S. Miniaturizing Whole-Cell Evolution: Novel Biological and Technical Perspectives. 36th Rabensteiner Kolleg. 23.05-02.06.2018, Pottenstein, Germany.
POSTER PRESENTATIONS
- Zoheir A.E. and Rabe K.S. RGB-S Reporter: A Novel Multi-Stress Whole-Cell Biosensor. Retreat 2018 of the Biointerfaces International Graduate School (BIF-IGS). 12-14.06.2018, Rothenburg odT, Germany.
- Zoheir A.E., Niemeyer C.M. and Rabe K.S. RGB-S Reporter: A Novel Multi-Stress Whole-Cell Biosensor. Annual meeting 2018 of the Association for General and Applied Microbiology (VAAM). 15-18.04.2018, Wolfsburg, Germany.
Featured Research 2022
Wide Press Coverage to the MESIF Chip
Press release of KIT and many other media platforms
Click here to read the paper
(Cover design: Ahmed Zoheir)
Featured Research 2021
Front Cover of "SMALL" Features the "evo.S" Chip
Volume 17, Issue 10 - March 11, 2021
Click here to read the abstract
(Cover design: Ahmed Zoheir)


